The perfect outbreak...for showcasing the value of sequencing
Pseudomonas outbreak brings genomics to the forefront
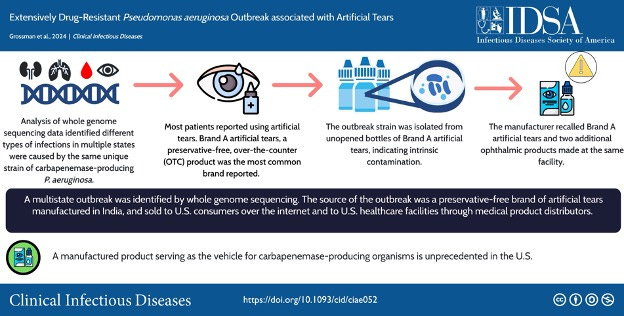
I got glasses when I was in second grade and probably could have used them a few years earlier. By the time I opted for LASIK as an adult, my contact lens prescription was -7.0, which meant that even with an alarm clock with two-inch-tall letters, I still had to squint to read the clock on my nightstand. Overall, LASIK has been well worth it, but one lingering side effect is dry eyes. As a result, I am an avid consumer of artificial tears, particularly in the winter. This personal background, along with my fascination with emerging infections and genomics, made the 2022–2023 carbapenemase-producing Pseudomonas aeruginosa (CRPA) outbreak especially interesting.
After learning more about the outbreak, I realized that, beyond its tragic story—at least four people died within 30 days of the first specimen collection, and at least four had an eye removed—it serves as a poster child for the power of pathogen genomics to make connections that no other technique can. In this article, I’ll walk through 5 different ways that this outbreak shows how sequencing can provide linkages across disciplines, organization types, time, space, and humans and animals.
Initial inklings of an outbreak
The outbreak is chronicled in a publication by Grossman et al. (2024)1. We learn that initially, healthcare facility clusters of CRPA were identified in three different states, which appeared unrelated until sequencing analysis revealed otherwise. This demonstrates sequencing’s ability to link cases across geography and jurisdictions.
Most (n = 26) cases confirmed prior to November 2022 belonged to 3 healthcare facility clusters in 3 states, which had been investigated as individual facility outbreaks before they were genetically linked by WGS analysis.
- Grossman et al. (2024)
Once the connection was made, investigators from multiple agencies collaborated to conduct epidemiologic investigations, product traceback, and testing, ultimately linking the outbreak to EzriCare brand artificial tears. Testing by the FDA identified positive products, and the sequences from these products were compared to human isolates, confirming the link. This highlights sequencing’s ability to link product samples to human isolates.
Healthcare and Academic investigations
While the investigation was ongoing within public health agencies, researchers and infection control professionals at the University of Pittsburgh Medical Center–Presbyterian Hospital were routinely conducting whole genome sequencing on potentially healthcare-associated Pseudomonas aeruginosa isolates2. Using information shared by the CDC about the sequence type and specific carbapenemase genes, as well as three example sequences, the Pittsburgh team identified two patients with isolates matching the outbreak pattern and alerted public health officials. This underscores sequencing’s role in connecting public health agencies with clinical and academic organizations.
In April 2023, after reviewing updates on the VIM-GES-CRPA artificial tears outbreak from the CDC including information about the ST of the outbreak strain and associated AMR genes, we retrospectively examined our WGS database for ST1203 P aeruginosa isolates and evaluated these for the presence of blaVIM-80 and blaGES-9. We identified 2 isolates from patient 1 and 1 isolate from patient 2 that belonged to ST1203 and encoded both carbapenemase genes at 100% identity.
- Sundermann et al. (2024)
Animal Health investigations
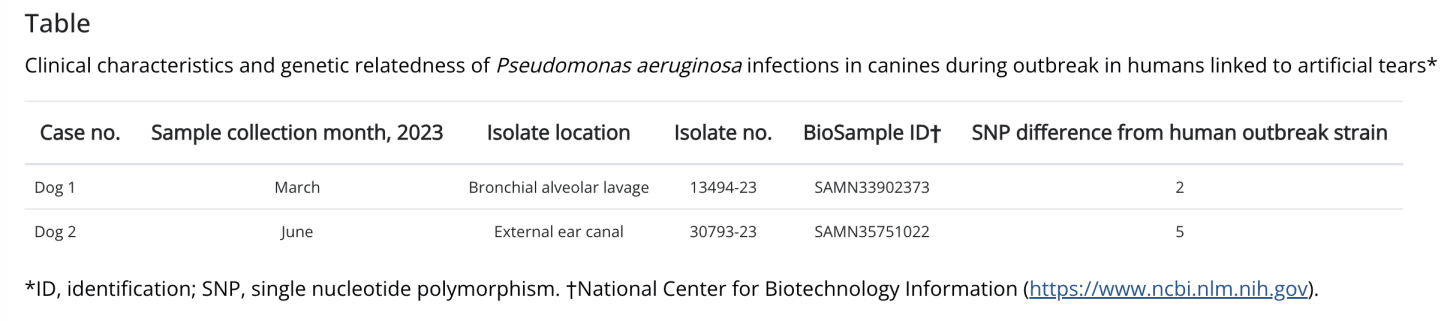
While the outbreak investigation was ongoing in the human sphere, the New Jersey Department of Health was notified of CRPA isolates from two different dogs treated at a single veterinary hospital in New Jersey. Their story is outlined in a recent publication by Price et al. (2024)3. Clinical specimens from the dogs were sent to the clinical microbiology laboratory of the PennVet Diagnostic Laboratory, University of Pennsylvania (Philadelphia, PA, USA) where they underwent culture, AST, and sequencing. Sequences were uploaded to NCBI and included in the NCBI Pathogen Detection (see this article for more about NCBI PDP)
The New Jersey Department of Health conducted a through epidemiologic investigation of both dogs as well as the veterinary hospital, and were able to provide infection prevention and control recommendations to the hospital facility. No direct link was found between the dogs, their owners, and the recalled product. This highlights one of the challenges of organisms like Pseudomonas that can be carried and spread asymptomatically, making it difficult to track the source of later cases that may have passed through several hosts. This detection in companion animals highlights a need for expanded surveillance and research on the interaction of carbapenemase-producing organisms across humans and their pets. This shows sequencing’s capacity to link across the human and animal health spheres.

Ongoing surveillance
Though the contaminated artificial tears were identified and recalled in February of 2023, isolates matching the characteristic outbreak pattern continue to be identified. Sequencing data is critical to be able to monitor the outbreak over time, and the NCBI Pathogen Detection Project (PDP) is a great way to be able to track - for example this query will return all Pseudomonas aeruginosa in the NCBI PDP that contain the blaVIM-80 and blaGES-9 genes. Of course, this only detects sequences that have been uploaded to NCBI and pass quality control metrics so it’s not comprehensive, but it’s a great resource to work with.
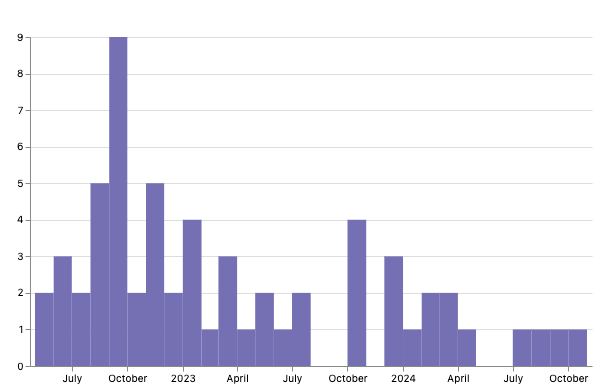
From the timeline below you can see sequences by collection date, and you can see the most recent collection date is October 2024. Detecting these isolates allows public health investigation into whether the new cases are linked to continued use of recalled product, or likely linked to onward person-to-person or environmental transmission. Being able to connect cases that occurred before the outbreak was identified with isolates from the outbreak as well as isolates that continue to be seen after the recall is a critical strength of sequencing data. This shows sequencing’s capacity to link across time.
Conclusions
This outbreak demonstrates the power of genomics as a bridge across multiple dimensions: between public health agencies and academic medical centers, between human and animal health, and across time. Much like music is often hailed as a universal language, sequencing can be seen as a universal language that connects different scientific disciplines and organizations.
The outbreak also highlights two key challenges in increasing the utility of sequencing for public health investigations. One is the technical challenge of increasing sequencing capacity, and the second is the need to build networks and data-sharing systems to ensure that those outside governmental public health agencies can fully contribute. Without incorporating data and input from clinical, academic, and veterinary sequencers, the system will fall short of its potential. Encouraging robust, timely, and open sequence data sharing is crucial to unlocking the full power of emerging sequencing technologies.
This outbreak serves as a powerful example because it clearly shows sequencing’s ability to link across:
Geography and jurisdictions
Product samples and human isolates
Public health agencies and clinical/academic organizations
Human and animal health spheres
Time
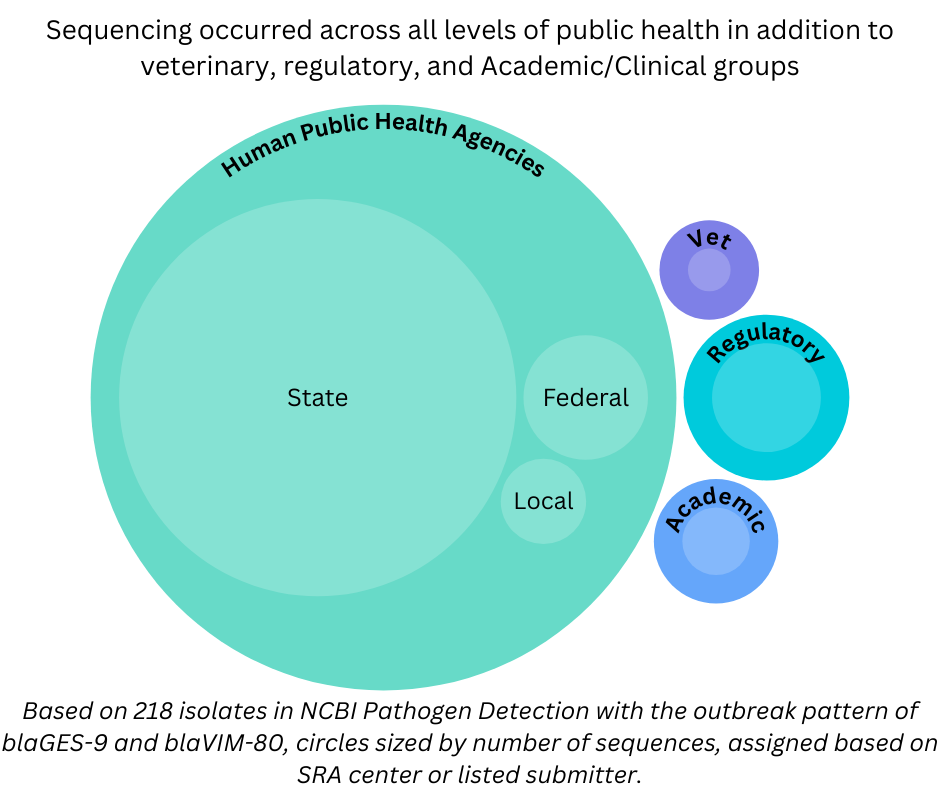
Within public health agencies, the ability to quickly link across different levels—from county public health labs to federal agencies—is another key strength. Further connecting these to results from veterinary, regulatory, and academic/clinical submitters ensures the most comprehensive surveillance system.
I have many favorite outbreaks (mostly ones I’ve worked on), but this one stands out because it so clearly demonstrates genomics' potential to transform public health. We should all strive to work more closely together in pursuit of improved health for all—humans and dogs alike.
References
Marissa K Grossman, Danielle A Rankin, Meghan Maloney, Richard A Stanton, Paige Gable, Valerie A Stevens, Thomas Ewing, Katharine Saunders, Sarah Kogut, Elizabeth Nazarian, Sandeep Bhaurla, Jehan Mephors, Joshua Mongillo, Susan Stonehocker, Jeanette Prignano, Nickolas Valencia, Argentina Charles, Kiara McNamara, William A Fritsch, Shannon Ruelle, Carrie Ann Plucinski, Lynn E Sosa, Belinda Ostrowsky, D Cal Ham, Maroya S Walters, for the Multistate Pseudomonas Outbreak Investigation Group , Extensively Drug-Resistant Pseudomonas aeruginosa Outbreak Associated With Artificial Tears, Clinical Infectious Diseases, Volume 79, Issue 1, 15 July 2024, Pages 6–14, https://doi.org/10.1093/cid/ciae052
Alexander J Sundermann, Vatsala Rangachar Srinivasa, Emma G Mills, Marissa P Griffith, Kady D Waggle, Ashley M Ayres, Lora Pless, Graham M Snyder, Lee H Harrison, Daria Van Tyne, Two Artificial Tears Outbreak-Associated Cases of Extensively Drug-Resistant Pseudomonas aeruginosa Detected Through Whole Genome Sequencing–Based Surveillance, The Journal of Infectious Diseases, Volume 229, Issue 2, 15 February 2024, Pages 517–521, https://doi.org/10.1093/infdis/jiad318
Price, E. R., McDermott, D., Sherman, A., Kelley, L., Mehr, J., Greeley, R....Cole, S. D. (2024). Canine Multidrug-Resistant Pseudomonas aeruginosa Cases Linked to Human Artificial Tears–Related Outbreak. Emerging Infectious Diseases, 30(12), 2689-2691. https://doi.org/10.3201/eid3012.240085.