Turning sequencing into surveillance
Who, where, what, when, and why of samples
One of the key decisions to be made when building genomic surveillance and epidemiology capacity is what samples should be sequenced. It’s important to bring all the stakeholders together because the needs of different groups can lead to very different conclusions on what samples should be sequenced. More samples is often helpful, but a scattershot approach of just increasing volume with no regard to representativeness or intended purpose of the data is often counterproductive, giving impressive numbers and terabytes of data, but little actual value for driving public health policy and decision making.
Population level surveillance
At a population level, one goal for sampling is to establish representative data to monitor over time what is happening in the population of interest. You have to know what is normal to quickly flag what is weird! Sampling should be designed in a way that provides adequate representation across critical characteristics such as geographic, race, sex, age, etc. The importance of this is easy to grasp when you think about extremes - would a wastewater surveillance system that only collected samples from elementary schools in one corner of a state be able to be used to describe the epidemiology of that disease in the state as a whole? Clearly not!
No surveillance system will be perfect and free from bias, but it’s important to design your system to be as good as possible for answering the questions you need it to answer, and ensure that you’re aware of any biases when you perform analyses. The image below of the surveillance cycle shows the steps walking through from data collection through to public health action. Importantly, the system evaluation piece allows you to review how well your surveillance system is meeting the goals and needs from your original system design process.
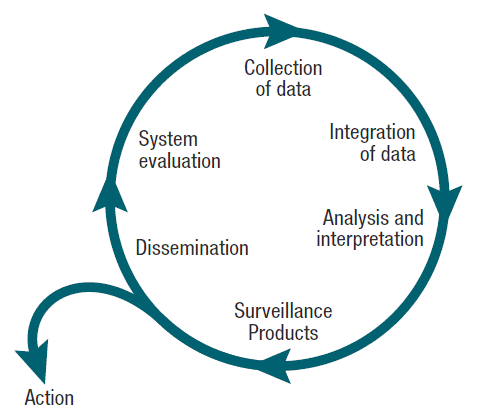
This intentional design and ongoing systematic nature of the sampling is what distinguishes a sequencing program from a genomic surveillance program. Importantly, It’s not just about collecting a nice representative(ish) dataset, but it’s about all the steps that come afterwards in the surveillance cycle - the analysis, interpretation, sharing, and action taken based upon the data. The action may be short-term such as recalling a food item, or longer-term such as changing policies for how compounding pharmacies are regulated.
The reason for collecting, analyzing, and disseminating information on a disease is to control that disease. Collection and analysis should not be allowed to consume resources if action does not follow.” - William Foege
Population-level sequencing in the COVID-19 pandemic
The COVID-19 pandemic led to an explosion of sequencing, and as of February 2024, there are over 16 million genomes available in the GISAID database - pretty wild! One effort to to better track sequences in public repositories that were from randomly selected vs. targeted specimens was the use of “baseline surveillance” tagging. The CDC asked sequencing laboratories to tag their randomly selected specimens in a specific manner in the databases which allows them to be identified by anyone conducting analyses. This helps reduce noise in the dataset by removing 50 samples from one facility outbreak, or thousands of samples from one hospital network that is only sequencing hospitalized patients. This is an improvement at a national scale, but still difficult to ascertain how representative the data is because of limited metadata in the sequence repositories.
In Washington State (where I was working at the time), we built a network of sentinel laboratories to try and improve representativeness of submissions statewide. A recent analysis by Hanna Oltean, et. al1 found that the sentinel laboratories program did improve representativeness overall in multiple categories (age, death from COVID-19, outbreak association, long-term care facility–affiliated status, and geographic coverage). This step of analyzing how well your surveillance system is doing is so important both for updating your surveillance system, and ensuring you’re drawing the correct conclusions from your data.
Hospitals: surveillance in special populations
There have been a couple of great papers recently covering ongoing genomic surveillance program within hospital settings. The first paper is from Stribling et. al2 at a US military hospital system which uncovered a decades-long P. aeruginosa cluster using routine sequencing (both retrospective and prospective). One key point they make is regarding the importance of surveillance sequencing rather than just picking ‘concerning’ isolates. From a laboratory surveillance perspective, selectively sequencing high-risk or unusual isolates is a good way to track emerging genes or discover new mechanisms of resistance, but from an infection-control and epidemiology perspective, broader surveillance is critical. This is why it’s important when designing sampling strategies to have clear communication across lab, epidemiology, infection prevention, etc. to create a system that meets the goals and needs of all data users.
Here, routine surveillance of MDR pathogens across the MHS was critical in uncovering a decades-long P. aeruginosa epidemic cluster. With traditional approaches alone20, comparable outbreaks may avoid detection due to the sporadic nature of patient infections, scattering of cases throughout the hospital, and changing antibiotic susceptibility profiles. Furthermore, because of budget constraints, surveillance programs focusing on “high-risk” isolates carrying select resistance genes (e.g. ESBLs, carbapenemases)4–6, would also fail to detect this ST-621 outbreak clone.
-William Stribling et. al
The second one is by Nathan Raabe et. al.3 and examines a multi-year multi-species outbreak of Enterobacterales associated with New Delhi metallo-β-lactamase (NDM) encoding plasmids in a hospital. This hospital had instituted routine WGS for selected high-priority healthcare-associated infections in 2021 (through a program called EDS-HAT4. By combining traditional infection prevention and control activities with the WGS data, they were able to detect this protracted outbreak. Importantly, many genomic surveillance systems are not designed to detect plasmid-mediated outbreaks which occur across multiple species like this one. Acknowledging and designing systems to accommodate the true complexity of the combination of bacterial transmission and horizontal plasmid transfer is critical to ensuring that we are not missing clusters and outbreaks because of limitations in our surveillance system design.
Another key point of the paper from the surveillance design perspective is the additional use of active surveillance. Whereas routine surveillance relies on isolates primarily collected for clinical diagnostic purposes, active surveillance is done in response to a detected cluster and involves proactively swabbing patients to look for more cases.
Fifty-three rounds of active surveillance were performed around 13 index patients, ranging from one to 17 rounds per patient (median, three rounds). Of 1,198 individuals meeting criteria for active surveillance, 913 (76·2%) had a peri-anal swab obtained. Among these, 2 (0·2%) were positive for the outbreak NDM-5-encoding plasmid; one additional case of outbreak NDM-5 plasmid carriage was identified during active surveillance around patients with non-outbreak NDM-1 enzyme carriage. Three additional patients were identified while performing routine active surveillance for carbapenem-resistant organisms
Raabe et. al.
The combination of routine plus active surveillance, and long and short-read sequencing creates a really comprehensive picture of what was occurring in the facility. These types of outbreaks are likely occurring in facilities routinely, but are not being detected. This type of comprehensive and integrated approach to genomic surveillance in a healthcare setting with direct connection to informing infection prevention and control is an excellent model.
Wrapping it up
In summary, getting the most out of sequencing data is about far more than getting as many sequences as possible. It is critical to first tease out the needs of the end users - do the labs need a certain number of isolates that meet specific criteria to validate an assay? Do the epidemiologists want broad population surveillance to track trends over time? Are there grant metrics that you have to meet? Okay, that last one is a bit tongue-in-cheek, but if it’s your reality, it’s something you have to work within.
Once these have been addressed and agreed upon, sampling plans can be created that allow the most benefit within the budget available. Genomic surveillance is a powerful tool as the two outbreak examples show, and getting it right requires strong cross-functional collaboration, but the benefits are well worth the investment of time and resources.
References
Oltean HN, Allen KJ, Frisbie L, et al. Sentinel Surveillance System Implementation and Evaluation for SARS-CoV-2 Genomic Data, Washington, USA, 2020–2021. Emerging Infectious Diseases. 2023;29(2):242-251. doi:10.3201/eid2902.221482.
Stribling William, Hall Lindsey R., Powell Aubrey, Harless Casey, Martin Melissa J., Corey Brendan W., Snesrud Erik, Ong Ana, Maybank Rosslyn, Stam Jason, Bartlett Katie, Jones Brendan T., Preston Lan N., Lane Katherine F., Thompson Bernadette, Young Lynn M., Kwak Yoon I., Barsoumian Alice E., Markelz Ana-Elizabeth, Kiley John L., Cybulski Robert J., Bennett Jason W., Mc Gann Patrick T., Lebreton Francois (2024) Detecting, mapping, and suppressing the spread of a decade-long Pseudomonas aeruginosa nosocomial outbreak with genomics eLife 13:RP9318 https://doi.org/10.7554/eLife.93181.1
Nathan J. Raabe, Abby L. Valek, Marissa P. Griffith, Emma Mills, Kady Waggle, Vatsala Rangachar Srinivasa, Ashley M. Ayres, Claire Bradford, Hannah M. Creager, Lora L. Pless, Alexander J. Sundermann, Daria Van Tyne, Graham M. Snyder, Lee H. Harrison, Real-Time Genomic Epidemiologic Investigation of a Multispecies Plasmid-Associated Hospital Outbreak of NDM-5-Producing Enterobacterales Infections, International Journal of Infectious Diseases, 2024, ISSN 1201-9712, https://doi.org/10.1016/j.ijid.2024.02.014.
Sundermann, A. J., Chen, J., Kumar, P., Ayres, A. M., Cho, S. T., Ezeonwuka, C., ... & Harrison, L. H. (2022). Whole-genome sequencing surveillance and machine learning of the electronic health record for enhanced healthcare outbreak detection. Clinical Infectious Diseases, 75(3), 476-482. https://doi.org/10.1093/cid/ciab946


